Introduction
Obesity is a chronic disease state caused by excessive accumulation of body fat. The Korean Society for the Study of Obesity defines the criteria for pre-obesity or overweight as a body mass index (BMI) of 23 kg/m2 or higher, and the criteria for obesity as a BMI of 25 kg/m2 or higher (KSSO, 2024). Obesity occurs when an individual's energy intake exceeds energy expenditure, leading to increased body fat. It involves a complex interplay of various risk factors, including dietary habits, lifestyle, age, genetic factors, and socioeconomic factors. Among the various mechanisms, genetic factors account for 10–20% of the cause (KSSO, 2024). To date, human genome research has identified numerous genetic loci associated with obesity. These genetic factors provide crucial clues for understanding how genetics influence the onset and progression of obesity in individuals.
Various genes associated with obesity are being studied. Representative examples include the FTO (Fat Mass and Obesity-associated) and MC4R (Melanocortin-4 Receptor) genes, which regulate food intake and energy homeostasis, and have been shown to be associated with BMI (Dougkas et al., 2013). These genes were primarily reported to be associated with obesity in studies involving predominantly Caucasian populations (Wu et al., 2010). Additionally, genes such as BDNF, which recognize satiety or regulate appetite, exist. Research on these genes primarily focuses on controlling obesity onset through dietary regulation (Unger et al., 2007).
Recently, polygenic risk scores (PRS) have been reported to examine genetic contributions to specific diseases. This method measures the cumulative contribution of individual single nucleotide polymorphisms (SNPs) present in a specific person (Oh et al., 2020). A PRS is a score that considers all genetically associated SNP variants potentially relevant to predicting an outcome. SNPs generally represent single nucleotide pair variations in the reference genome and play a central role in calculating PRS. PRS is not merely a tool for assessing the risk of specific diseases; it also provides crucial information for summarizing the cumulative genetic susceptibility to disease related metabolic traits (Jaenisch & Bird, 2003). Understanding genetic factors in obesity is essential not only for obesity itself but also for reducing the onset of chronic diseases induced by obesity and developing prevention strategies. Genomic research for obesity prevention and management is rapidly being adopted in both clinical practice and direct-to-consumer (DTC) testing fields. In the market, genetic markers discovered through Western studies are already actively offered as solutions for obesity management (Oh, 2019). In Korea, research is progressing by collecting genetic information, lifestyle habits, diet, clinical data, and follow-up results, primarily based on large-scale national cohorts of individuals aged 40 and above, to analyze the gene-environment (lifestyle)-disease pathways that actually function in Koreans (Kwak & Shin, 2023). Specifically, large-scale datasets reflecting the genetic characteristics and lifestyle habits of Koreans are being constructed through standardized repeated surveys and follow-up studies, the establishment of genomic databases via biobanks, linkage with external data (insurance, mortality, cancer registries, environmental exposures, etc.), and Genome-wide association studies (GWAS)/PRS/interaction analyses.
While diverse domestic research is underway, commercial services in Korea do not exclusively incorporate genes identified through Korean Genome and Epidemiology Study (KoGES) data. In other words, the rapid expansion and diversity of the domestic DTC market are currently not sufficiently supported by scientific evidence grounded in Korean genetics. The realization of precision medicine necessitates the collection and analysis of large-scale healthcare information; in this context, research analyzing the genetic factors of obesity using healthcare big data is essential for establishing personalized prevention and treatment strategies. Therefore, a comparative analysis of genetic markers associated with the Korean population, utilizing both Western-based genetic markers and KoGES data, must be conducted as a priority. Accordingly, this study employed a scoping review to analyze research exploring obesity-related genetic factors from 2010 to 2025, based on KoGES data, with the aim of identifying the status of obesity genes validated in the Korean population.
Materials and Methods
Scoping reviews are appropriately utilized to analyze key points, types of evidence, and differences between studies within large research datasets (Colquhoun et al., 2014). This study conducted an analysis following the five steps of the scoping review framework by Arksey & O'Malley (2005) to identify trends in research aiming to determine genetic risk factors for obesity. The detailed research content for each step is as follows.
First, deriving the research question involves linking and clarifying the research purpose and the research question. For scoping reviews, it is recommended to set broad questions to aid in establishing a search strategy and to facilitate a broad understanding of the research domain. Therefore, this study set the following research question to systematically review the trends in studies analyzing the association between obesity and genetic factors based on KoGES data and to identify which Korean genes influence obesity.
The literature search was conducted from March 1, 2025, to October 31, 2025, and investigated papers examining the association between genetic factors and obesity from studies conducted from 2010 to 2025. To minimize potential search omissions during the data retrieval process, a total of five academic search databases were utilized. For domestic literature, databases (RISS, KISS, and DBpia) were searched, while international literature was retrieved from databases (PubMed and ScienceDirect). The search function was designed to identify studies that simultaneously met three sub-criteria. The search function was designed to find results satisfying three sub-criteria simultaneously. These sub-criteria included search terms for obesity, obesity-related biomarkers, obesity-related genetic factors, and KoGES. The search strategy across the five databases utilized Boolean operators (AND, OR) to combine keywords. The specific search string was constructed as follows: (“BMI” OR “Abdominal obesity” OR “Obesity” OR “Waist Circumference”) AND (“KoGES” OR “Ansan/Anseong” OR “HEXA”) AND (“Gene” OR “SNP” OR “Single nucleotide polymorphism” OR “GRS” OR “Genetic risk score” OR “GWAS” OR “PRS” OR “Polygenic risk score” OR “DNA methylation”).
The first procedure in the study selection process involved screening titles and abstracts of studies relevant to the topic. Subsequently, the full-text articles were reviewed to select the final study documents. The criteria for selecting and excluding study documents considered during the review are as follows.
-
(1) Study population: Research involving Korean adults aged 19 years or older
-
(2) Data source: Research utilizing data from the KoGES
-
(3) Genetic factors: Studies that performed GWAS or used SNPs identified through GWAS or PRS as primary variables
-
(4) Obesity indicators: Studies investigating associations with obesity-related traits such as obesity, weight, BMI, abdominal obesity, waist circumference
-
(5) Study type: Original research articles published in domestic or international academic journals
-
(6) Language: Studies published in English
-
(1) Duplicate studies: Literature retrieved from different databases and identified as duplicates
-
(2) Unclear variable definitions: Studies where the definitions or measurement criteria for obesity and obesity-related indicators were not clearly described
-
(3) Inappropriate study type: Non-original research such as non-peer-reviewed literature, dissertations, or conference abstracts
-
(4) Unavailable full text: Cases where the full text of the paper could not be obtained
The retrieved records were managed using bibliographic software (EndNote) to remove duplicates. Titles and abstracts were subsequently screened to assess eligibility according to the predefined selection criteria. When eligibility could not be determined based on the title and abstract alone, the full text was reviewed for inclusion.
The final step 5 is the stage of understanding the meaning of the research results. Scoping reviews differ most significantly from systematic reviews in that they provide comprehensive data on research in a relevant field and do not evaluate study quality to draw general conclusions. This study selected and recorded research according to the scoping review methodology, ultimately analyzing and summarizing the research findings to derive a consensus conclusion.
Results
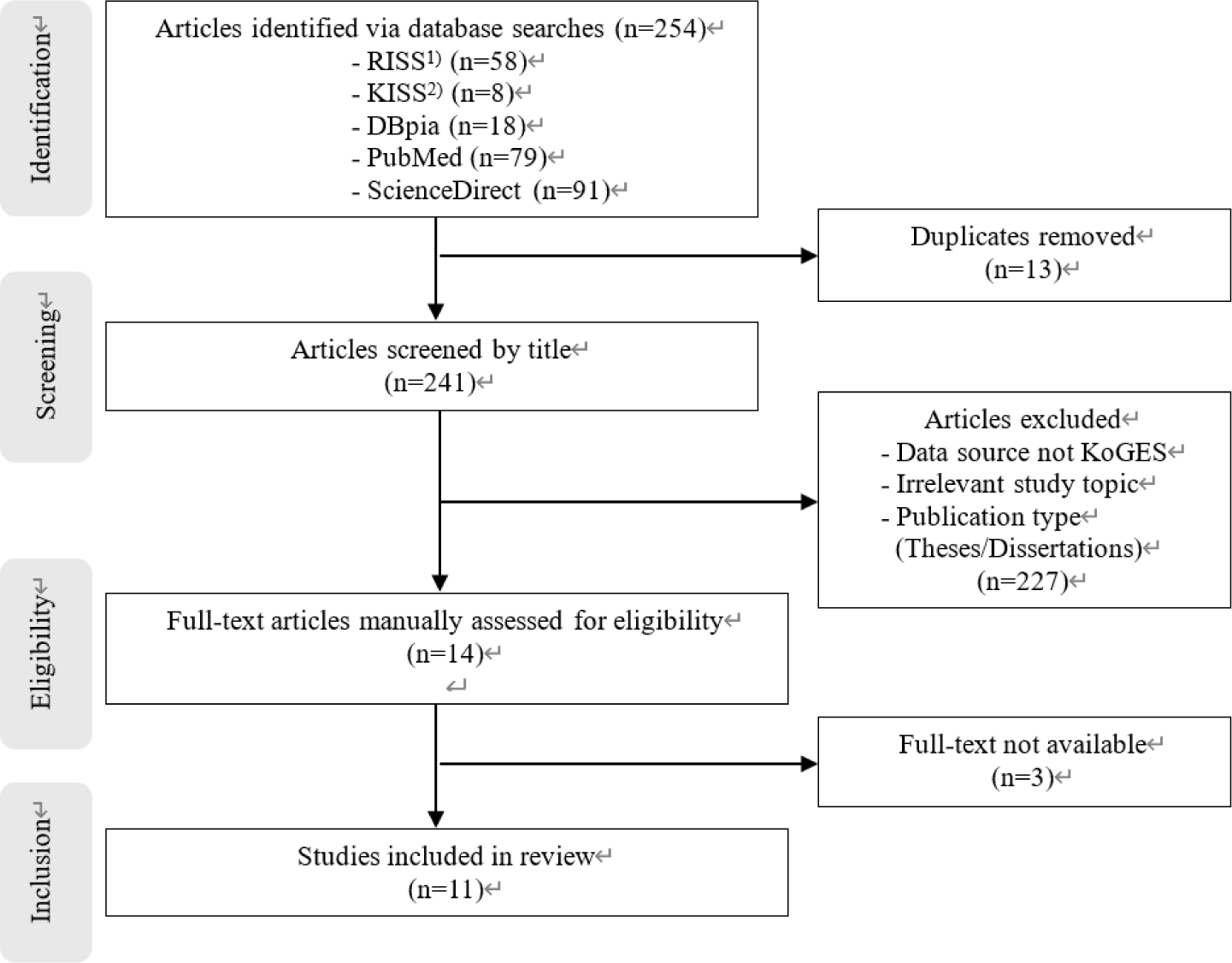
Searching five academic databases (RISS, KISS, DBpia, PubMed, ScienceDirect) using the English keywords “BMI”, “Abdominal obesity”, “Obesity”, “Waist Circumference”, “KoGES”, “Ansan/Anseong”, “HEXA”, “Gene”, “SNP”, “Single nucleotide polymorphism”, “GRS”, “Genetic risk score”, “GWAS”, “PRS”, “Polygenic risk score” and “DNA methylation”. Among studies published over the 15-year period from 2010 to 2025, a total of 254 papers analyzed genetic factors related to metabolic syndrome using KoGES data. First, we excluded 13 duplicated articles found across academic databases. Subsequently, 230 articles were excluded based on selection and exclusion criteria, resulting in a final inclusion of 11 articles in this study (Fig. 1).
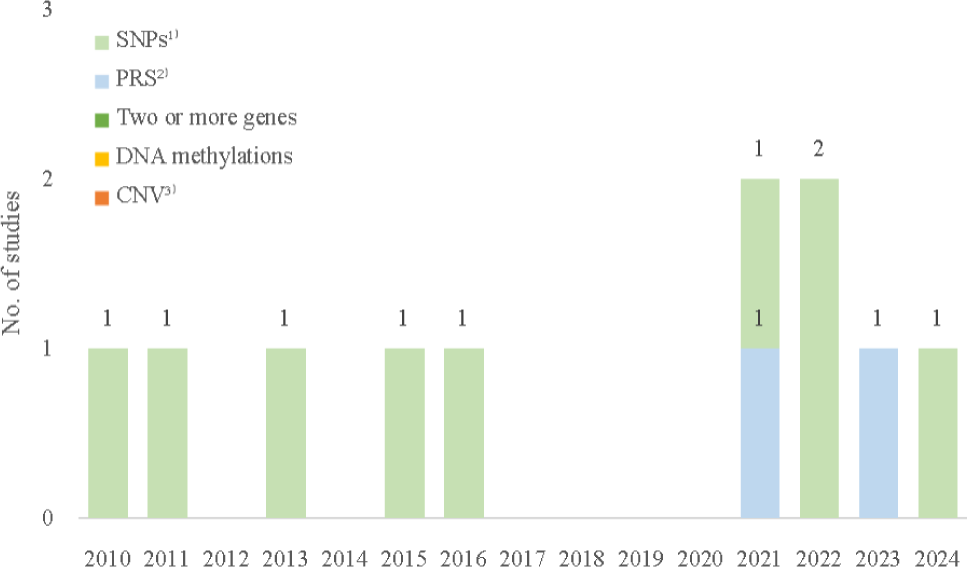
The results of analyzing studies published over the past 15 years that explored obesity-related genetic risk factors in Korean adults using KoGES data, categorized by publication year, are presented in Fig. 2.
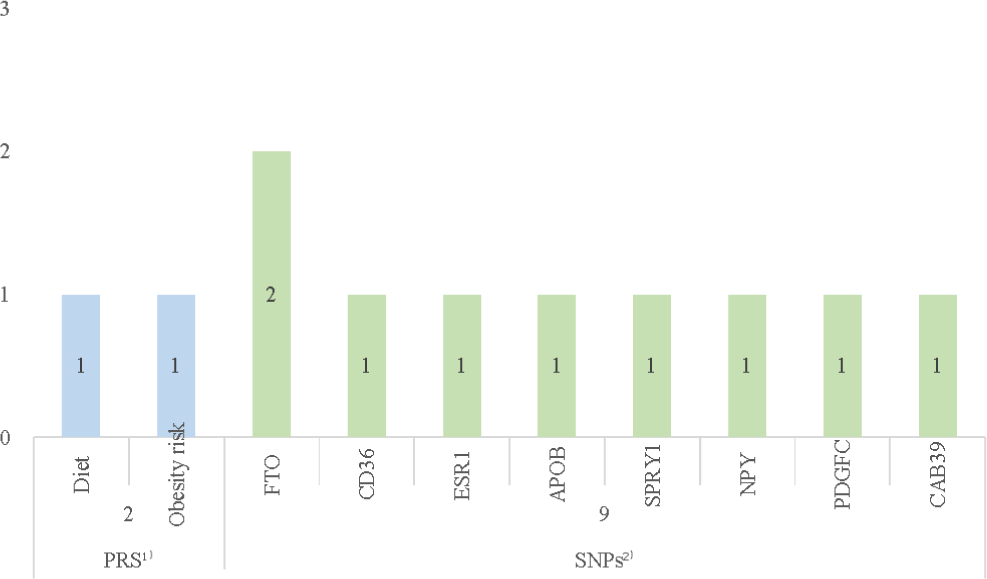
A total of 11 studies analyzed genetic risk factors for obesity, including 9 SNPs and 2 papers on PRS (Fig. 3).
The scope of research has expanded from simple SNP analysis to more sophisticated genetic prediction models. The key characteristics and summaries of the 11 papers ultimately selected for this study are presented in Table 1.
| No. | Study design | Studies | Genetic approach | Outcome (s) | Data source (s) | Sample size (n) | Conclusions |
|---|---|---|---|---|---|---|---|
| 1 | Cross-Sectional study | Lee et al. (2021) | PRS1) | BMI2) |
KARE3) CAVAS4) HEXA5) |
8,444 9,300 17,350 |
PRS and aPRS were significantly associated with BMI and obesity, but no significant interaction was observed with total calorie or macronutrient intake. |
| 2 | Cross-sectional & Longitudinal study | Yoon & Cho (2023) | PRS | Obesity Metabolic traits |
KARE CAVAS HEXA |
13,504 (longitudinal study 5,400) |
The developed PRS effectively predicted obesity and related metabolic diseases. Longitudinal analysis showed that a higher PRS was associated with an increased incidence of dyslipidemia and hypo-HDL cholesterolemia. |
| 3 | Cross-sectional | Lee et al. (2010) |
FTO rs9939609 |
BMI | KARE | 8,842 | Common FTO variants were associated with BMI and overweight in adults. A significant interaction was observed between rs9939609 and physical activity, whereas no association with dietary fat intake was found in adults. |
| 4 | Case-control | Doo & Kim (2011) |
ESR1 rs1884051 |
BMI | KARE | 3,039 | The ESR1 rs1884051 polymorphism was significantly associated with obesity-related variables in men. This association was modified by total energy and plant protein intake; specifically, the minor T allele was associated with lower BMI in the high plant protein intake group. |
| 5 | Cross-sectional study | Jin et al. (2013) |
SPRY1 rs923982 |
%BF6) %AbF7) BMI WHR8) |
KARE | 3,013 | SPRY1 gene polymorphisms and the TGCC haplotype were significantly associated with increased body fat percentage, abdominal fat, and osteoporosis risk in Korean women. |
| 6 | Case-control | Doo et al. (2015) |
APOB rs1469513 |
BMI obesity | KoGES | 6,470 | The association between the APOB rs1469513 polymorphism and obesity was significantly modified by dietary fat intake; specifically, high fat intake increased obesity risk in minor G allele carriers. |
| 7 | Cross-sectional | Kim et al. (2016) |
NPY rs16149 |
BMI WC9) VAT10) |
KARE | 1,468 | Common NPY polymorphisms were not directly associated with obesity but showed significant interactions with psychosocial stress on BMI, waist circumference, and visceral adipose tissue (VAT). |
| 8 | Cross-sectional | Choi (2021) |
CD36 rs1527479 |
Dietary intake | KARE | 6,619 | The CD36 polymorphism was associated with cruciferous vegetable intake in obese males; those with the risk genotype consumed significantly less vegetables, while no association was found with fat intake. |
| 9 | Cross-sectional | Goh & Choi (2022) |
FTO rs1121980 |
BMI Dietary intake |
KARE | 6,262 | The FTO rs1121980 variation was associated with a preference for high-fat foods (e.g., coffee creamer, snacks), and these preferences varied by sex and BMI. |
| 10 | Longitudinal | Lee et al. (2024) |
PDGFC rs4691380 TREH rs2276064 |
Longitudinal BMI change | KARE | 3,074 | Identified specific genetic variants (e.g., PDGFC, TREH) significantly associated with long-term BMI changes and obesity risk in Korean adults. |
| 11 | Cross-sectional | Kwon et al. (2022) |
CAB39 rs6722579 CPQ rs59465035 |
Abdominal obesity | KoGES | 50,808 | Specific genetic variants interacted with nutrient intake to influence obesity risk; CAB39 variant increased abdominal obesity risk with high fat intake, while CPQ variant decreased risk with high vitamin C intake. |
The FTO, expressed in the hypothalamus to regulate appetite and energy balance, is the most influential obesity-related gene identified via GWAS. Its SNPs are consistently associated with increased BMI and obesity risk across diverse populations (Lee et al., 2010; Goh & Choi, 2022). Lee et al. (2010) confirmed in a study of 8,842 Korean adults that FTO gene variants (rs9939973, rs9939609) showed a significant positive correlation with obesity indicators such as BMI and waist circumference and were also significantly associated with fasting insulin and HOMA-IR levels. Logistic regression analysis revealed that carriers of FTO variants (particularly rs9939609) had a significantly higher risk of overweight, approximately 1.16 times greater than non-carriers. Goh & Choi (2022) analyzed the association between the rs1121980 variant of the FTO gene and dietary intake patterns in 6,262 individuals. The FTO gene variant did not show a significant association with macronutrient intake itself, such as total energy intake, carbohydrate, protein, or fat intake. However, among obese women, carriers of the T allele consumed significantly more coffee cream than non-carriers. Among obese men, T allele carriers tended to consume more snacks. Thus, it was confirmed that the FTO gene variant may contribute to obesity risk not by simply increasing overall food intake, but by increasing preference for foods high in fat or energy density, such as coffee creamers and snacks.
CD36 facilitates fatty acid transport and serves as a fat taste receptor. Variations in the CD36 gene modulate fat taste sensitivity, potentially increasing high-fat food intake and obesity risk. Choi (2021) analyzed the association between the rs1527479 polymorphism of the CD36 gene and dietary intake patterns in 3,194 men and 3,425 women. The analysis revealed differences in dietary intake based on obesity status. Among obese men, the group with the minor allele homozygous genotype (CC genotype) reported significantly lower intake of cruciferous vegetables (Brassicaceae) compared to the group with the major allele heterozygous genotype (TT/CT).
The ESR1 gene encodes estrogen receptor alpha (ERα) and plays a pivotal role in regulating energy metabolism and fat distribution. Specifically, it is known to suppress obesity by regulating adipose tissue function and fat distribution through the modulation of alpha-2A adrenergic receptor–mediated lipolytic signaling in subcutaneous fat (Doo & Kim, 2011). Doo & Kim (2011) analyzed the effects of the rs1884051 polymorphism in ESR1 on obesity indicators and its interaction with dietary intake in 3,039 middle-aged Korean men. The results showed that males carrying the minor allele (T allele) of rs1884051 had significantly lower body weight, BMI, body fat percentage, and waist-to-hip ratio compared to those carrying the major allele (C allele), and their obesity risk was also 0.79 times lower. In groups with low total energy intake or high plant protein intake, BMI was significantly lower in individuals carrying the T allele. This suggests that the ESR1 variant acts as a factor lowering obesity risk in Korean men and can be modulated by dietary factors (plant protein intake).
The APOB gene encodes a major component of LDL cholesterol involved in lipid transport. It is associated with weight gain by directly influencing energy storage and metabolic processes (Doo et al., 2015). Doo et al. (2015) analyzed the effect of the rs1469513 polymorphism in the APOB gene on obesity in 6,470 individuals aged 40–59. The group carrying the minor allele (G allele) of this variant showed significantly higher weight and BMI compared to those carrying the major allele (A allele). Furthermore, it was confirmed that the obesity risk associated with the APOB gene is critically influenced by fat intake: in the high-fat diet group, G allele carriers had a 1.31-fold higher risk of obesity compared to A allele carriers. This indicates that the APOB variant is a potential obesity risk factor, and a high-fat diet amplifies the obesity-inducing effect of this gene.
The SPRY1 gene regulates mesenchymal stem cell differentiation, functioning as a key molecular switch that suppresses adipogenesis and fat production (Jin et al., 2013). Jin et al. (2013) analyzed the effect of SPRY1 gene variants on obesity traits in 3,013 Korean women. The study found that the rs923982 variant showed a significant association with body fat percentage (%BF) and abdominal fat percentage (%AbF). Specifically, haplotype analysis revealed that women carrying the TGCC type (rs300555, rs10518414, rs923982, rs12650992) had significantly higher body fat percentage (p=0.0087) and abdominal fat percentage (p=0.047) compared to those without this haplotype. These results indicate that SPRY1 gene variants act as a genetic factor promoting fat accumulation and increasing obesity risk in Korean women.
NPY is a neurotransmitter that stimulates appetite and fat production. Notably, stress-induced glucocorticoids promote NPY secretion, which accelerates abdominal fat accumulation, serving as a key link between chronic stress and obesity (Kim et al., 2016). Kim et al. (2016) analyzed the interaction between NPY gene variants and stress on BMI, waist circumference, and visceral adipose tissue (VAT) in 1,468 individuals. The NPY gene variant alone showed no significant association with obesity indicators. However, when analyzed in conjunction with stress, significant associations were confirmed with BMI, waist circumference, and VAT. Notably, homozygotes for the rs16149 allele (AA) had an obesity prevalence of only 23.53% in the low-stress group, but this surged to 72.22% in the high-stress group. This suggests that the NPY gene variant acts as a factor increasing the risk of abdominal and visceral obesity under stressful conditions.
PDGFC promotes adipose tissue expansion via adipogenesis and cell proliferation (Lee et al., 2024). Lee et al. (2024) conducted a 14-year longitudinal analysis of 3,074 Koreans to examine genetic effects on long-term BMI changes. The study revealed that the PDGFC rs4691380 variant was significantly associated with long-term BMI increases in both men and women. Unlike cross-sectional evidence that links genes to obesity status at a single time point, this longitudinal finding highlights that PDGFC influences the trajectory of obesity progression by inducing sustained weight gain over time.
CAB39 is an upstream regulator of the AMP-activated protein kinase pathway, which regulates cellular energy metabolism, and is known to be involved in regulating adipocyte differentiation and lipolysis (Kwon et al., 2022).
Kwon et al. (2022) analyzed the interaction between the intake levels of 19 nutrients and genetic variants on obesity and abdominal obesity in 50,808 individuals. The study found that the rs6722579 variant in the CAB39 gene interacted with dietary fat intake. In the group with fat intake higher than the recommended level, carriers of the minor allele for this variant had a significantly higher risk of abdominal obesity, approximately 3.73 times greater than non-carriers. This suggests that a high-fat diet is an environmental factor that promotes abdominal fat accumulation in carriers of the CAB39 variant.
Because the influence of a single genetic variant is limited, PRS that combine obesity-related genetic variants to assess an individual's genetic susceptibility have been identified in numerous studies. Lee et al. (2021) analyzed the interaction between obesity PRS and total calorie and macronutrient intake in a study of 35,094 individuals across three cohorts. In all three cohorts, individuals with high PRS had significantly higher BMI regardless of dietary habits. However, the interaction between PRS and total calorie intake was not statistically significant, and this pattern was consistently observed in analyses of macronutrient intake such as protein, fat, and carbohydrates. This indicates that genetic obesity risk and dietary intake contribute independently to obesity, and even in genetically susceptible individuals, there is no clear evidence of a particularly increased obesity risk due to calorie intake. The report did not mention representative genes contributing to the PRS. Meanwhile, Yoon et al. (2023) developed a Korean-specific BMI PRS using KoGES data. The PRS, developed by combining 53,341 genetic variants, was found to explain approximately 2.4% of BMI variation in Koreans. When applying this model, the high-risk group (top 25%) showed a statistically significant approximately twofold higher risk of obesity compared to the low-risk group (bottom 25%). The difference in obesity prevalence between the top 10% and bottom 10% of PRS scores was approximately 26.0%. These findings indicate that, despite the modest variance explained by the PRS, it retains utility for population-level risk stratification rather than individual-level prediction. In addition, the PRS was significantly associated with fasting insulin and triglyceride levels, suggesting a potential link with metabolic risk profiles related to obesity. Genetic markers closely associated with obesity reported in the study include FTO, SEC16 homolog B (SEC16B), brain-derived neurotrophic factor (BDNF), transmembrane protein 18 (TMEM18), and cyclic nucleotide-binding domain-containing protein 2 (CNBD2). Therefore, the developed PRS can be utilized as an effective indicator for screening and predicting obesity risk based on an individual's genetic predisposition.
The current status of obesity genetic testing services provided by major domestic medical institutions and DTC companies was examined (Table 2).
In accordance with relevant domestic laws and initial regulations (MOHW, 2016) regulations, the BMI-associated genes FTO, MC4R, and BDNF have been approved for analysis at officially reported genetic testing agencies. Company A, a medical institution-contracted testing agency, focused on evaluating energy consumption efficiency and fat metabolism capacity using β3-Adrenergic Receptor (ADRB3), Peroxisome Proliferator-Activated Receptor-γ (PPAR-γ), and Uncoupled Protein-1 (UCP-1) as key genes. Company B was found to comprehensively analyze BMI and body fat percentage risk, including not only metabolism-related genes but also markers based on GWAS such as FTO and MC4R. In contrast, DTC companies included services such as genetic analysis specialized for dietary behavior and precision nutrition analysis. Company C analyzed not only appetite-regulating genes like FTO and MC4R but also segmented behavioral factors such as obesity risk, abdominal obesity, and food cravings. Company D was confirmed to provide tailored services focusing on personalized nutrition management based on taste preferences and nutrient metabolism capabilities, incorporating lipid metabolism genes like Fatty Acid Desaturase 1 (FADS1) and Cholesteryl Ester Transfer Protein (CETP) alongside MC4R and BDNF. Genes utilized in obesity genetic testing services partially included those based on KoGES data, while most employed genetic markers associated with populations primarily from the US or Europe.
Discussion
This study conducted a scoping review to examine the current landscape of obesity-related genetic research in Korean populations. Findings from single-gene variant studies indicate that obesity risk is regulated through diverse biological pathways that extend beyond simple energy metabolism, including interactions with dietary factors and stress responses. For example, SPRY1 has been shown to contribute to fat accumulation by regulating adipocyte differentiation, while PDGFC has been identified as a persistent risk factor associated with long-term weight gain. Variants in the FTO gene have been linked to increased preferences for high-calorie foods, such as coffee creamers and sweets, whereas CD36 has been reported to contribute to dietary imbalances, including avoidance of vegetable intake. With respect to dietary interactions, APOB and CAB39 have been shown to increase obesity risk under high-fat dietary conditions, while ESR1 is associated with a reduced risk of obesity in the context of plant-based protein intake. Furthermore, NPY has been found to markedly increase the risk of abdominal obesity under conditions of elevated psychological stress.
A study examining the association between integrating genetic factors (PRS) and obesity reported that genetic risk and total calorie intake independently contribute to obesity. It also found that obesity risk varies depending on dietary control even among high-risk genetic groups (Lee et al., 2021). Another study demonstrated that a Korean-specific BMI PRS could identify individuals at approximately double the risk of obesity onset in high-risk groups and account for about 26.0% of the prevalence difference, showing its potential as an indicator for precision medicine (Yoon et al., 2023).
Genes with established associations with obesity based on the Korean genomes include FTO, CD36, ESR1, APOB, SPRY1, NPY, PDGFC, and CAB39. However, analysis of the current status of domestic obesity gene testing services reveals that they primarily explain obesity in conjunction with gene analysis of Hepatocyte nuclear factor 4 gamma (HNF4G), Copine 4 (CPNE4), Poliovirus Receptor Related Protein 3 (PVRL3), R-spondin 3 (RSPO3), and others, which were primarily selected based on results from large-scale GWAS conducted on Western populations.
However, studies evaluating whether numerous obesity-related SNPs reported in populations of European descent function identically in East Asian populations have reported that some loci discovered in Europeans were not replicated with the same direction or effect size (Lu & Loos, 2013). According to Nakayama et al. (2014), the Regulatory associated protein of mTOR complex 1 gene (RPTOR) polymorphism, known as a Western obesity gene, showed no significant association with obesity traits in Japanese adults. There are reports that FTO variants, including rs8050136 and rs9939609, showed no significant association with BMI, obesity, or body fat mass in Chinese Han populations (Li et al., 2008), and some meta-analysis results have reported only weak associations in China or certain East Asian groups (Peng et al., 2011). Kim et al. (2007) also pointed out that while PPAR-γ variants are used as obesity prediction markers in Western populations to lower BMI or improve insulin sensitivity, in Koreans, the variant frequency itself is low, and the association with obesity traits is insignificant. Furthermore, it has been reported that the Proprotein convertase subtilisin/kexin type 1 (PCSK1) gene variant, which was shown to increase obesity risk in studies on Europeans, was not detected in Chinese populations (Qi et al., 2010). Therefore, applying genetic variants identified in other ethnic groups directly to Koreans may result in reduced predictive accuracy.
This study has several limitations. While obesity-related genes were reported with statistical significance in the studies included in this review, most were based on observational study designs. Therefore, the potential influence of residual confounding factors cannot be ruled out, necessitating caution in interpreting the results. Consequently, the interaction findings between the genes presented in this review and obesity should be understood as evidence at the level of association rather than causation. In particular, lifestyle factors such as diet, physical activity, and stress not only influence obesity onset but may also be altered by obesity status, necessitating consideration of reverse causation. Nevertheless, the systematic compilation and identification of obesity-related genes validated in the Korean population in this study is expected to serve as foundational data for developing Korean-specific genetic analysis services and personalized health management strategies in the future.
In conclusion, while numerous genetic variants have been associated with obesity, the KoGES-based studies reviewed in this article have most consistently identified genes such as FTO, CD36, ESR1, APOB, SPRY1, NPY, PDGFC, and CAB39. Despite the diverse mechanisms of obesity-related genes being elucidated through international research, applying these findings directly to Koreans has limitations. Domestic epidemiological studies on how these genes interact with Korean-specific lifestyle habits to produce actual phenotypes are relatively scarce compared to the pace of commercialization. Therefore, for current domestic obesity genetic testing services to move beyond simply using markers based on Western populations and utilize markers optimized for Koreans, a validation process using large-scale data from Korean populations, such as the KoGES is necessary. Furthermore, in-depth research to elucidate the interactions between genetic and environmental factors appears to be continuously required. If such research is conducted, it is anticipated that the implementation of precision nutrition services tailored to Koreans will be possible through the integrated analysis of Korean-specific genetic variations and lifestyle habit data.
